DNA Sequencing
A molecular technique to reveal the arrangement of nucleotides in polynucleotide chains...
Second Generation
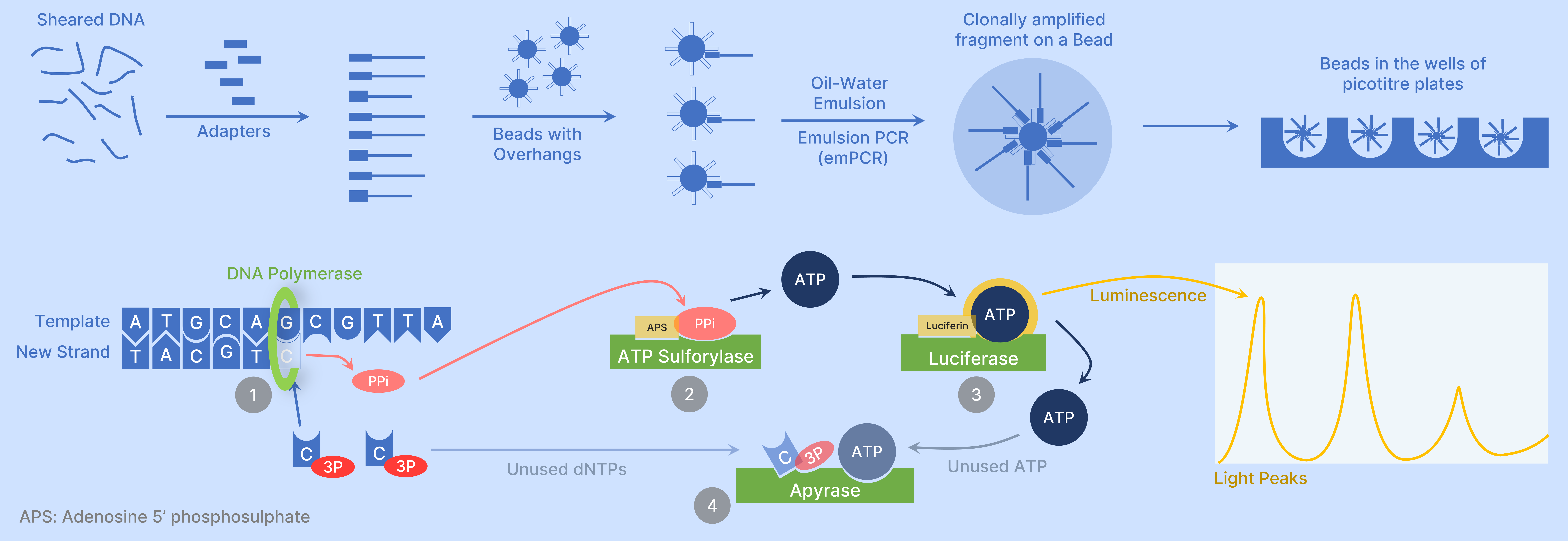
Pyrosequencing
- Principle described by Bertil Pettersson, Mathias Uhlen and Pål Nyren
- Sequence-by-Synthesis (SBS)
- Based on the detection of luminescence produced by the co-activity of 3 enzymes
- DNA Polymerase
- ATP Sulfurylase
- Luciferase
- Apyrase (Post-sequencing activity)
- Microfluidic variant developed by Jonathan Rothberg (454 Life Sciences)
- Output Format: SFF
- Read length: 300-500 bases
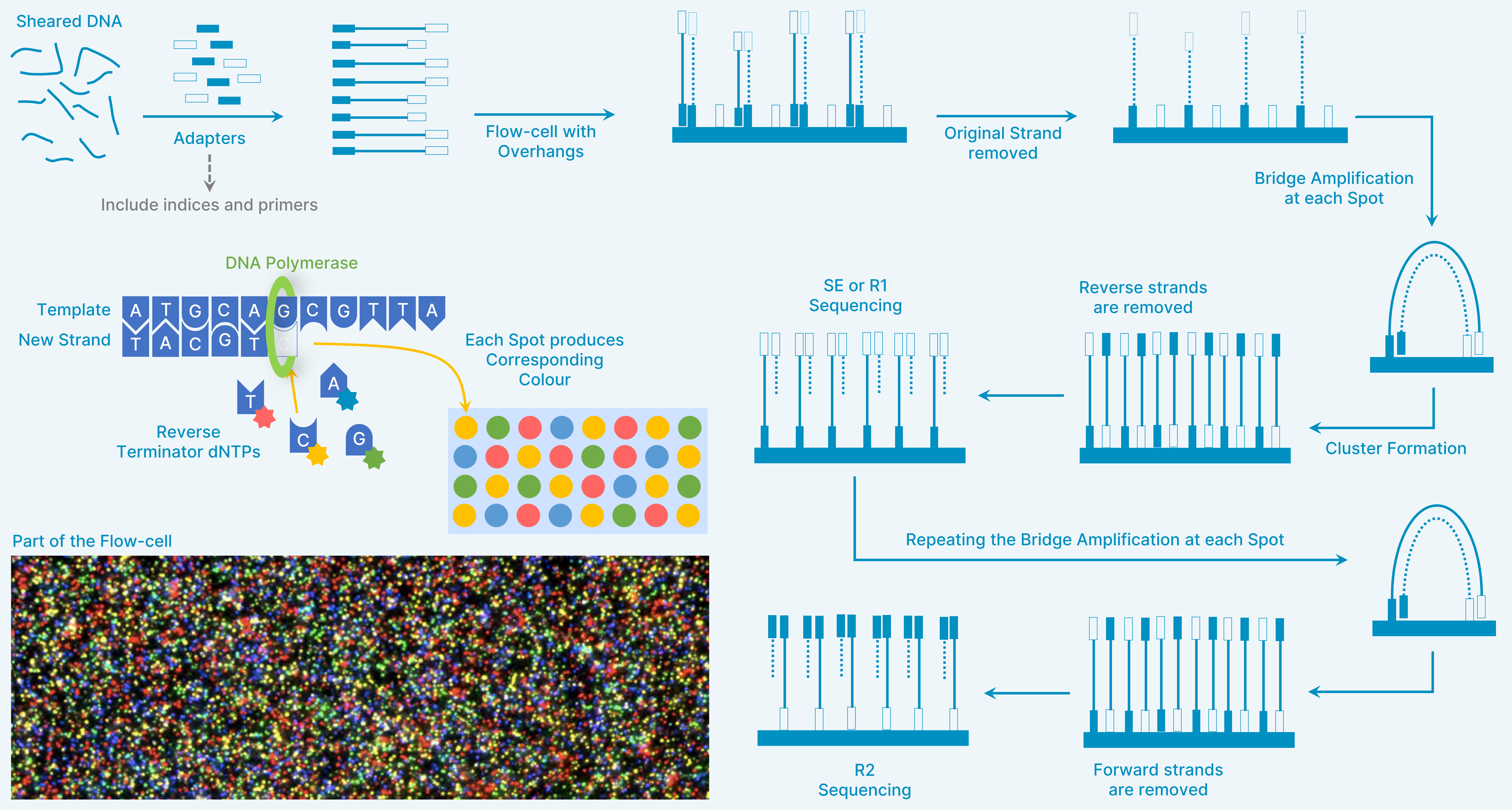
Short Read Sequencing (Illumina)
- Concept invented by Bruno Canard and Simon Sarfati (Pasteur Institute, Paris)
- Developed by Shankar Balasubramanian and David Klenerman (Solexa)
- Reversible dye-terminators
- Sequence-by-Synthesis (SBS)
- Genomic Library Preparation - Sonification or Tagmentation (Transposases) - 50 to 500 bp fragments
- Adapters have 3 parts - Adapter sequence, Index sequence, and Primer sequence
- Output format: fastq
- Read length: 35 to 300 bp
- Instruments: GA, GAII, MiSeq, HiSeq, NovaSeq, etc...
Third Generation
Single Molecule Sequencing (SMS) by Stephen Quake, commercialized by Helicos Biosciences - Similar to Illumina sequencing, excluding the bridge amplification step. First technology to sequence non-amplified DNA thus avoiding the amplification associated biases an errors. Slow and expensive
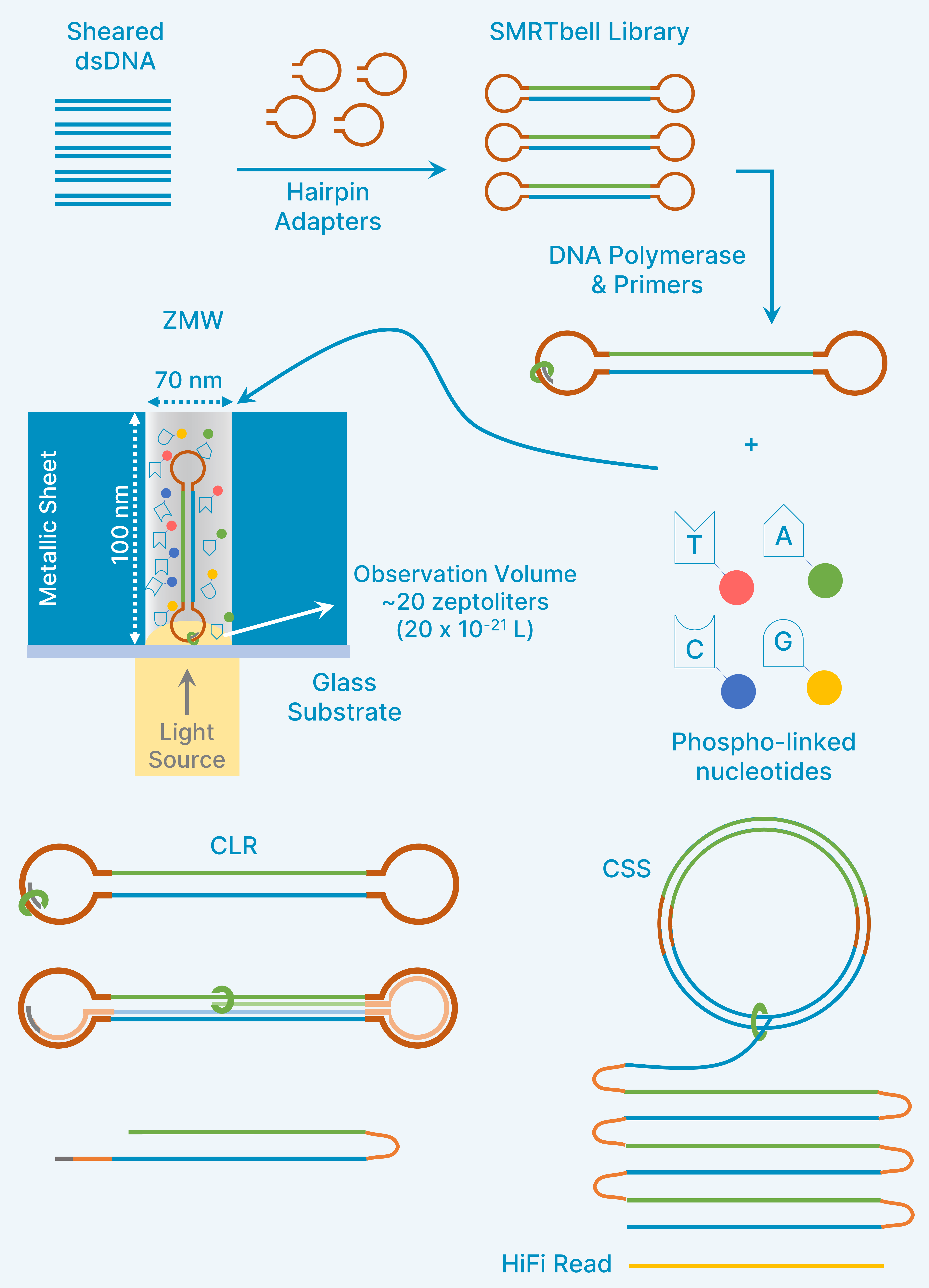
Single Molecule Real Time (SMRT) Sequencing (PacBio)
- Hairpin adaptors
- Phospholinked nucleotides with fluorophores
- SMRTbell library
- Zero-Mode Waveguides (ZMWs)
- Continuous Long Read Sequencing (CLR)
- Circular Consensus Sequencing (CSS)
- Average Read Length: 10,000 bp - 100,000 bp
- Instruments: RS, RS II, Sequel
- Output format: BAM
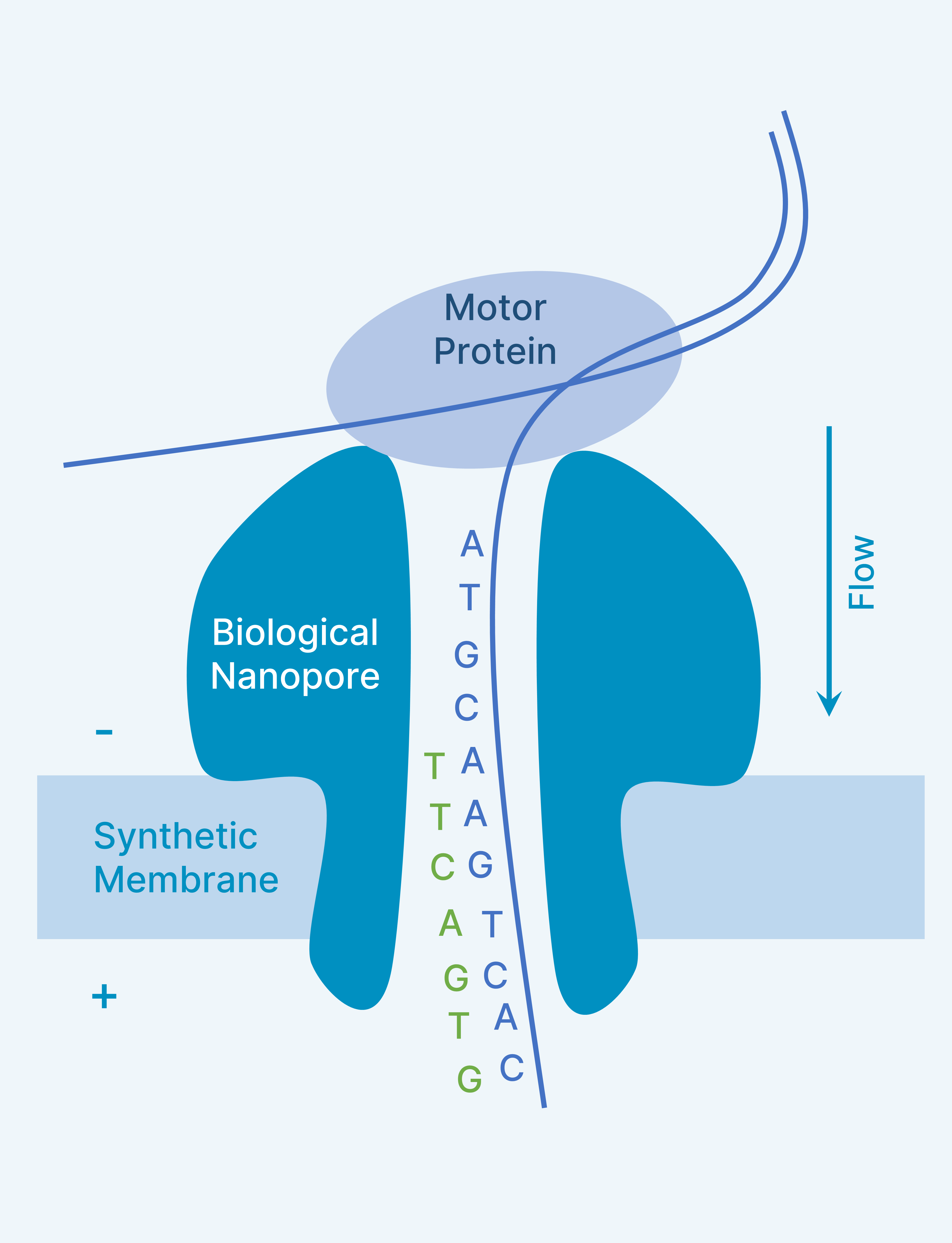
Nanopore Sequencing (Oxford)
- Idea proposed by David Deamer in 1989
- Hagan Bayley co-founded Oxford Nanopore in 2005
- Uses Biological Nanopores like α-hemolysin, MspA, CsgG
- Synthetic membranes
- Submerged in ionic solution
- Single strand from a dsDNA molecule is fed into the nanopore with the help of motor protein
- Variations in the ionic current while incorporating different nucleotides is captured
- Read Length of ~1 Mbp can be achieved
- Instruments: MinION, PromethION, etc...
- Output format: FAST5