Raw Reads QC
(Bacterial Genome Analysis Piplene)
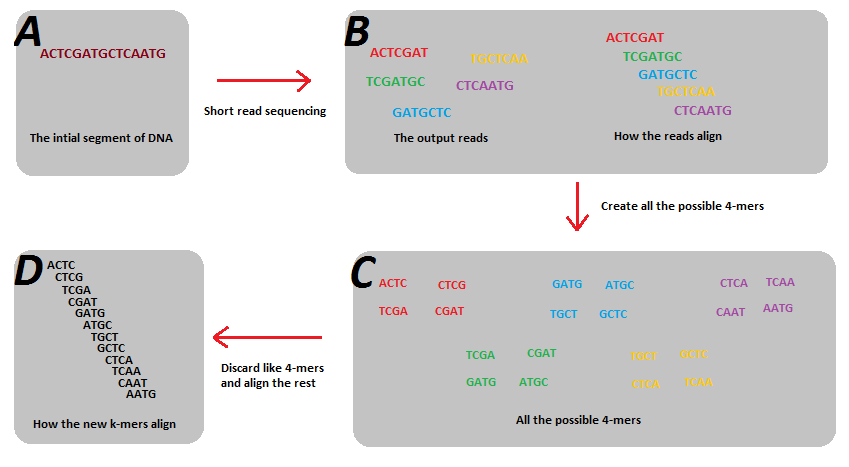
K-mers & De Bruijn Graphs
k-mers are substrings of length k. Example
Why k-mers are required
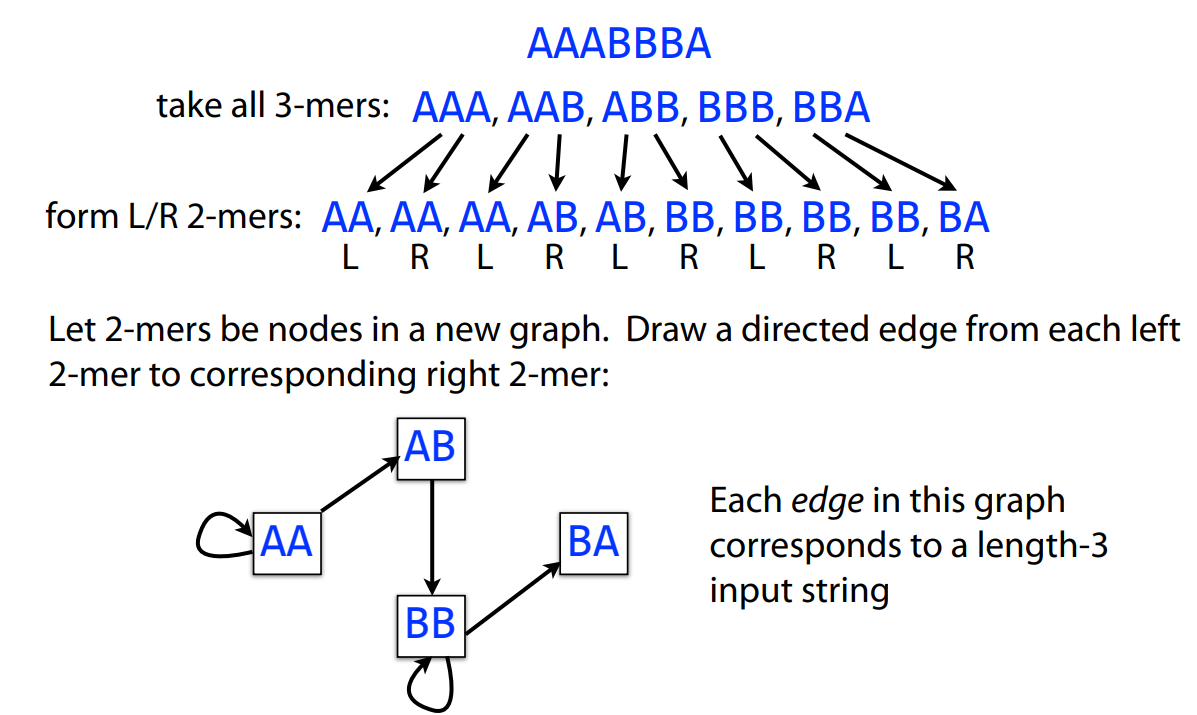
De Bruijn graph a directed graph representing overlaps between sequences of symbols
Example
A Quality Control Tool. Works on FASTQ, SAM and BAM files
-
Navigate to bgap directory and activate
qc$ cd Desktop/bgap $ conda deactivate $ conda activate qc -
Open FastQC GUI. Analyze and save the reports
$ fastqc - See the Basic statistics, Per base quality, Sequence length distribution, Overrepresented sequences and Adapter content sections
-
Run bbduk. Copied adapters file
$ mkdir bb_out $ cd bb_out $ bbduk.sh in1=../reads/a45_R1.fastq in2=../reads/a45_R2.fastq out1=a45_R1.fastq out2=a45_R2.fastq ref=adapters.fa k=23 mink=7 ktrim=r hdist=1 qtrim=r trimq=20 minlen=100 tpe tbo - Explanation:
-
Result
Input: 1741880 reads 436749684 bases. QTrimmed: 1522186 reads (87.39%) 103642774 bases (23.73%) KTrimmed: 376743 reads (21.63%) 13100754 bases (3.00%) Trimmed by overlap: 8692 reads (0.50%) 88862 bases (0.02%) Total Removed: 170588 reads (9.79%) 116832390 bases (26.75%) Result: 1571292 reads (90.21%) 319917294 bases (73.25%) -
Open FastQC GUI. Analyze and save the reports
$ fastqc
-
Run trimmomatic. Using BBDuk adapters file
$ mkdir trim_out $ cd trim_out $ trimmomatic PE -phred33 ../reads/a45_R1.fastq ../reads/a45_R2.fastq a45_R1_paired.fq.gz a45_R1_unpaired.fq.gz a45_R2_paired.fq.gz a45_R2_unpaired.fq.gz ILLUMINACLIP:../adapters.fa:2:30:10 SLIDINGWINDOW:4:20 MINLEN:100 - Input Read Pairs: 870940 Both Surviving: 599798 (68.87%) Forward Only Surviving: 160897 (18.47%) Reverse Only Surviving: 28249 (3.24%) Dropped: 81996 (9.41%)
-
Open FastQC GUI. Analyze and save the reports
$ fastqc